library(tidyverse) # 加载 tidyverse 包,提供数据科学常用工具
library(tidymodels) # 加载 tidymodels 包,提供构建和评估预测模型的函数
library(ISLR) # 加载 ISLR 包,包含用于统计学习的示例数据集
library(rpart.plot) # 加载 rpart.plot 包,用于绘制决策树图
library(vip) # 加载 vip 包,用于评估模型特征的重要性
load(
file = "D:/Document/0.Study R/0.R4DS/data/practice-data/pimadiabetes.rdata"
)
test <- pimadiabetes %>%
as_tibble() %>%
select(diabetes, everything())
pimadiabetes <- pimadiabetes %>%
as_tibble()
set.seed(123)
pima_split <- pimadiabetes %>%
initial_split(prop = 0.7)
pima_train <- training(pima_split)
pima_test <- testing(pima_split)48 随机森林
- 随机森林是一种集成算法(ensembnle),属于其中的袋装法(bagging),它将许多决策树模型的结果组合起来。
- 随机森林使用自助法进行重抽样,每次使用全部观测的2/3作为训练集,剩余的1/3作为测试集(这部分数据称为袋外数据),对袋外数据进行的预测叫做袋外预测或样本外预测。
- 随机森林支持分类、回归、生存分析等多种任务类型,且对数据要求不高,即使不进行预处理也没问题(但不能出现缺失值)。
- 随机森林是集合了多棵树的集成模型,它天生就容易过拟合。针对随机森林的调优思路就是限制模型的复杂度,从而防止模型过拟合,例如限制树的数量、节点样本数、分支样本数等,调优时只需要选择其中1~2个进行优化即可。
48.1 加载包
48.2 创建模型
- 使用
randomForest引擎创建随机森林模型。 - 设置
mtry参数为.cols(),表示随机森林分支时使用的预测变量数量为总预测变量数。 -
set_engine()中添加importance = TRUE参数,以保存相关变量的重要性信息,用于后续使用vip包可视化模型变量的重要性。
Random Forest Model Specification (classification)
Main Arguments:
mtry = .cols()
Engine-Specific Arguments:
importance = permutation
Computational engine: ranger # 训练模型
bagging_fit <- bagging_spec %>%
fit(diabetes ~ ., data = pima_train)
bagging_fitparsnip model object
Ranger result
Call:
ranger::ranger(x = maybe_data_frame(x), y = y, mtry = min_cols(~.cols(), x), importance = ~"permutation", num.threads = 1, verbose = FALSE, seed = sample.int(10^5, 1), probability = TRUE)
Type: Probability estimation
Number of trees: 500
Sample size: 537
Number of independent variables: 8
Mtry: 8
Target node size: 10
Variable importance mode: permutation
Splitrule: gini
OOB prediction error (Brier s.): 0.158174 # A tibble: 231 × 2
.pred_class diabetes
<fct> <fct>
1 neg neg
2 pos neg
3 pos pos
4 neg neg
5 pos neg
6 neg neg
7 neg neg
8 pos pos
9 neg neg
10 pos pos
# ℹ 221 more rows# 混淆矩阵
augment(bagging_fit, new_data = pima_test) %>%
conf_mat(truth = diabetes, estimate = .pred_class) Truth
Prediction pos neg
pos 126 31
neg 24 50# 准确率
augment(bagging_fit, new_data = pima_test) %>%
accuracy(truth = diabetes, estimate = .pred_class)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 accuracy binary 0.762# roc_auc
augment(bagging_fit, new_data = pima_test) %>%
roc_auc(truth = diabetes, .pred_pos)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 roc_auc binary 0.849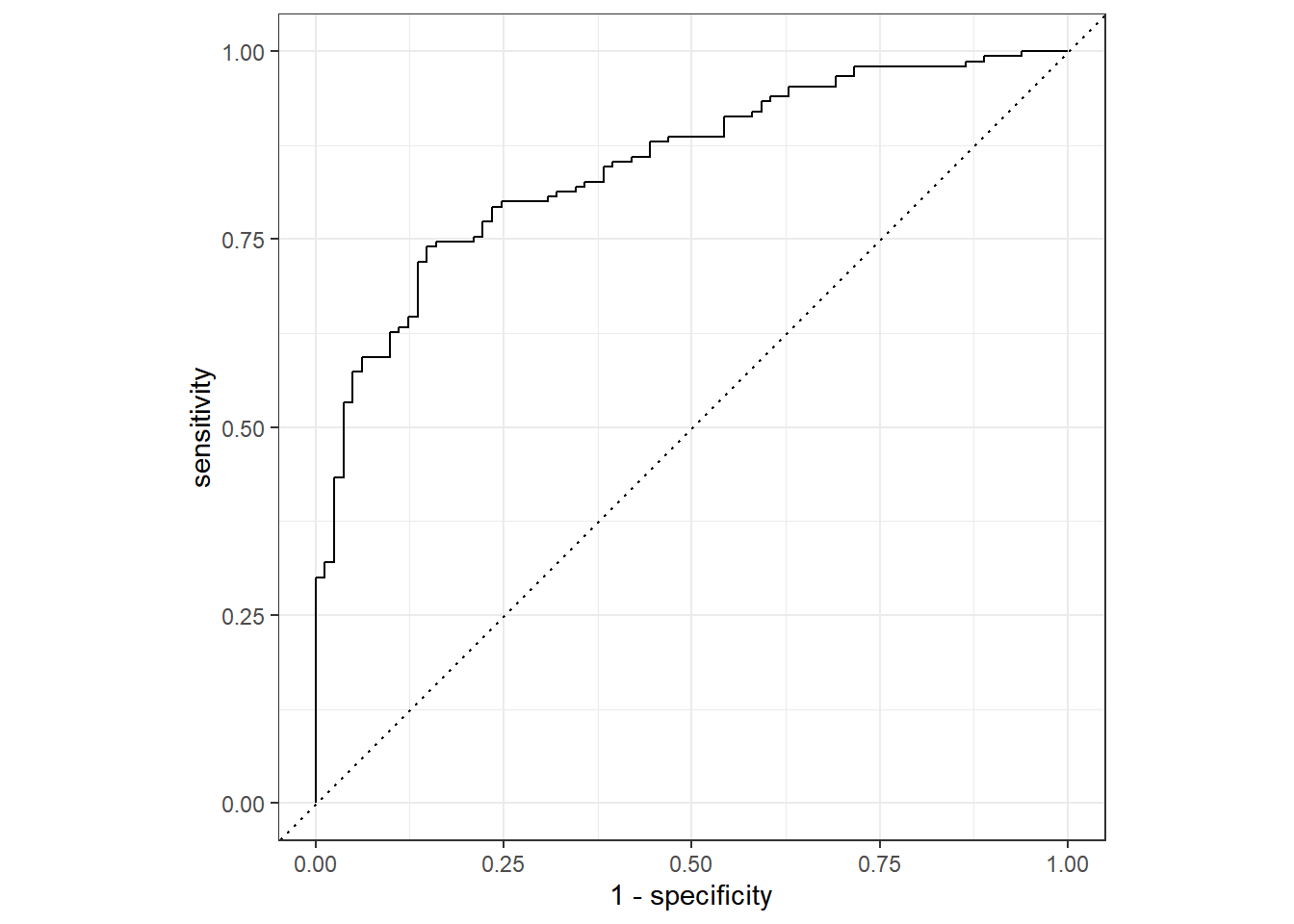
# 绘制变量重要性图形
vip(bagging_fit)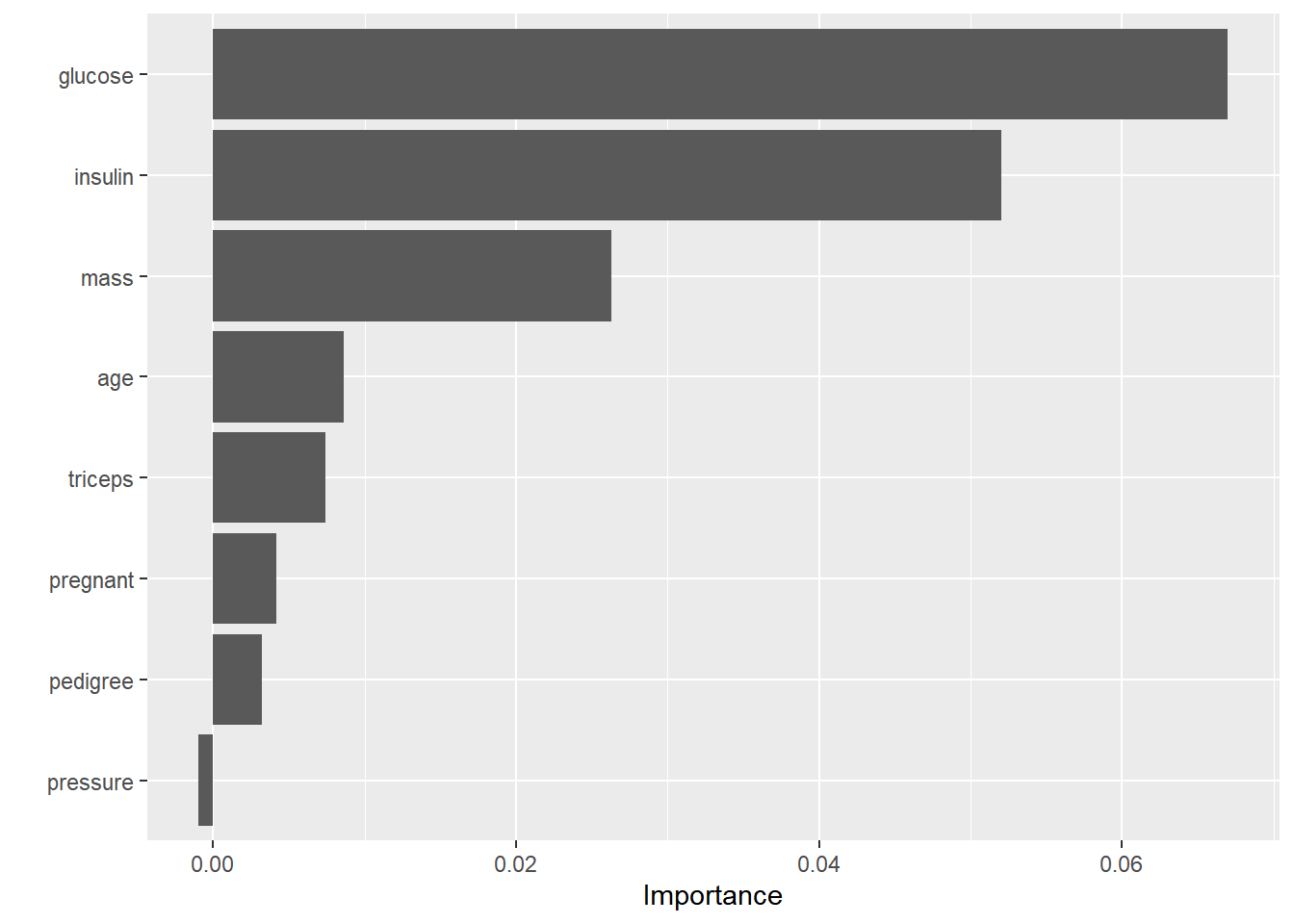
- roc_auc曲线显示,模型的AUC值为0.846 > 0.7,说明模型的预测能力较强。
- 变量重要性图形显示,
glucose、insulin和mass是随机森林模型的主要特征。
48.3 模型调优
随机森林模型中有三个可以优化的超参数,分别是:
-
mtry:随机森林分支时使用的预测变量数量。 -
trees:树的数量。 -
min_n:分支时需要的数据点的数量。
我们可以尝试调整这三个参数,以获得更好的模型性能。先用usemodels自动生成一个模型调优的过程看看。
library(usemodels)
use_ranger(
diabetes ~ .,
data = pima_train,
prefix = "ranger",
verbose = TRUE # 添加注释以解释代码含义
)ranger_recipe <-
recipe(formula = diabetes ~ ., data = pima_train)
ranger_spec <-
rand_forest(mtry = tune(), min_n = tune(), trees = 1000) %>%
set_mode("classification") %>%
set_engine("ranger")
ranger_workflow <-
workflow() %>%
add_recipe(ranger_recipe) %>%
add_model(ranger_spec)
set.seed(59134)
ranger_tune <-
tune_grid(ranger_workflow, resamples = stop("add your rsample object"), grid = stop("add number of candidate points"))下面进行超参数调优,基于树的模型对数据要求不严格,甚至可以不进行预处理。但为了保证过程的连贯性,我们还是统一使用workflow对象并添加recipes。
ranger_spec <- rand_forest(
mtry = tune(),
min_n = tune(),
trees = 1000
) %>%
set_mode("classification") %>%
set_engine("ranger", importance = "permutation")
ranger_recipe <-
recipe(formula = diabetes ~ ., data = pima_train)
ranger_wflow <- workflow() %>%
add_model(ranger_spec) %>%
add_recipe(ranger_recipe)
set.seed(1234)
ranger_folds <- vfold_cv(pima_train)
ranger_params <- ranger_wflow %>%
extract_parameter_set_dials() %>%
finalize(pima_train)
ranger_grid <- grid_regular(
ranger_params,
levels = 5
)
# metric_set <- metric_set(roc_auc, accuracy)
ranger_tune <- ranger_wflow %>%
tune_grid(
resamples = ranger_folds,
grid = ranger_grid,
# metrics = mretric_set,
control = control_grid(save_pred = TRUE)
)
show_best(ranger_tune)# A tibble: 5 × 8
mtry min_n .metric .estimator mean n std_err .config
<int> <int> <chr> <chr> <dbl> <int> <dbl> <chr>
1 1 40 roc_auc binary 0.847 10 0.0196 pre0_mod05_post0
2 3 30 roc_auc binary 0.847 10 0.0190 pre0_mod09_post0
3 3 40 roc_auc binary 0.846 10 0.0194 pre0_mod10_post0
4 3 21 roc_auc binary 0.846 10 0.0192 pre0_mod08_post0
5 1 30 roc_auc binary 0.845 10 0.0196 pre0_mod04_post0# 拟合最终模型
## 选择最优超参数
ranger_best <- select_best(ranger_tune, metric = "accuracy")
## 将最优参数应用至workflow
ranger_final_wflow <- ranger_wflow %>%
finalize_workflow(ranger_best)
ranger_final_wflow══ Workflow ════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: rand_forest()
── Preprocessor ────────────────────────────────────────────────────────────────
0 Recipe Steps
── Model ───────────────────────────────────────────────────────────────────────
Random Forest Model Specification (classification)
Main Arguments:
mtry = 7
trees = 1000
min_n = 40
Engine-Specific Arguments:
importance = permutation
Computational engine: ranger ## 拟合最终模型
ranger_final_fit <-
ranger_final_wflow %>%
last_fit(pima_split)
ranger_final_fit# Resampling results
# Manual resampling
# A tibble: 1 × 6
splits id .metrics .notes .predictions .workflow
<list> <chr> <list> <list> <list> <list>
1 <split [537/231]> train/test split <tibble> <tibble> <tibble> <workflow># 模型结果
ranger_final_fit %>%
collect_metrics()# A tibble: 3 × 4
.metric .estimator .estimate .config
<chr> <chr> <dbl> <chr>
1 accuracy binary 0.753 pre0_mod0_post0
2 roc_auc binary 0.847 pre0_mod0_post0
3 brier_class binary 0.154 pre0_mod0_post0# A tibble: 231 × 4
diabetes .pred_class .pred_pos .pred_neg
<fct> <fct> <dbl> <dbl>
1 neg neg 0.187 0.813
2 neg pos 0.606 0.394
3 pos pos 0.997 0.00258
4 neg neg 0.309 0.691
5 neg pos 0.635 0.365
6 neg neg 0.382 0.618
7 neg pos 0.552 0.448
8 pos pos 0.873 0.127
9 neg neg 0.253 0.747
10 pos pos 0.895 0.105
# ℹ 221 more rows## 混淆矩阵
conf_mat(final_augment, truth = diabetes, estimate = .pred_class) Truth
Prediction pos neg
pos 126 33
neg 24 48## roc_auc
roc_auc(final_augment, truth = diabetes, .pred_pos)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 roc_auc binary 0.847## 多个指标一同输出
ranger_metrics <- metric_set(accuracy, mcc, f_meas)
ranger_metrics(final_augment, truth = diabetes, estimate = .pred_class)# A tibble: 3 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 accuracy binary 0.753
2 mcc binary 0.446
3 f_meas binary 0.816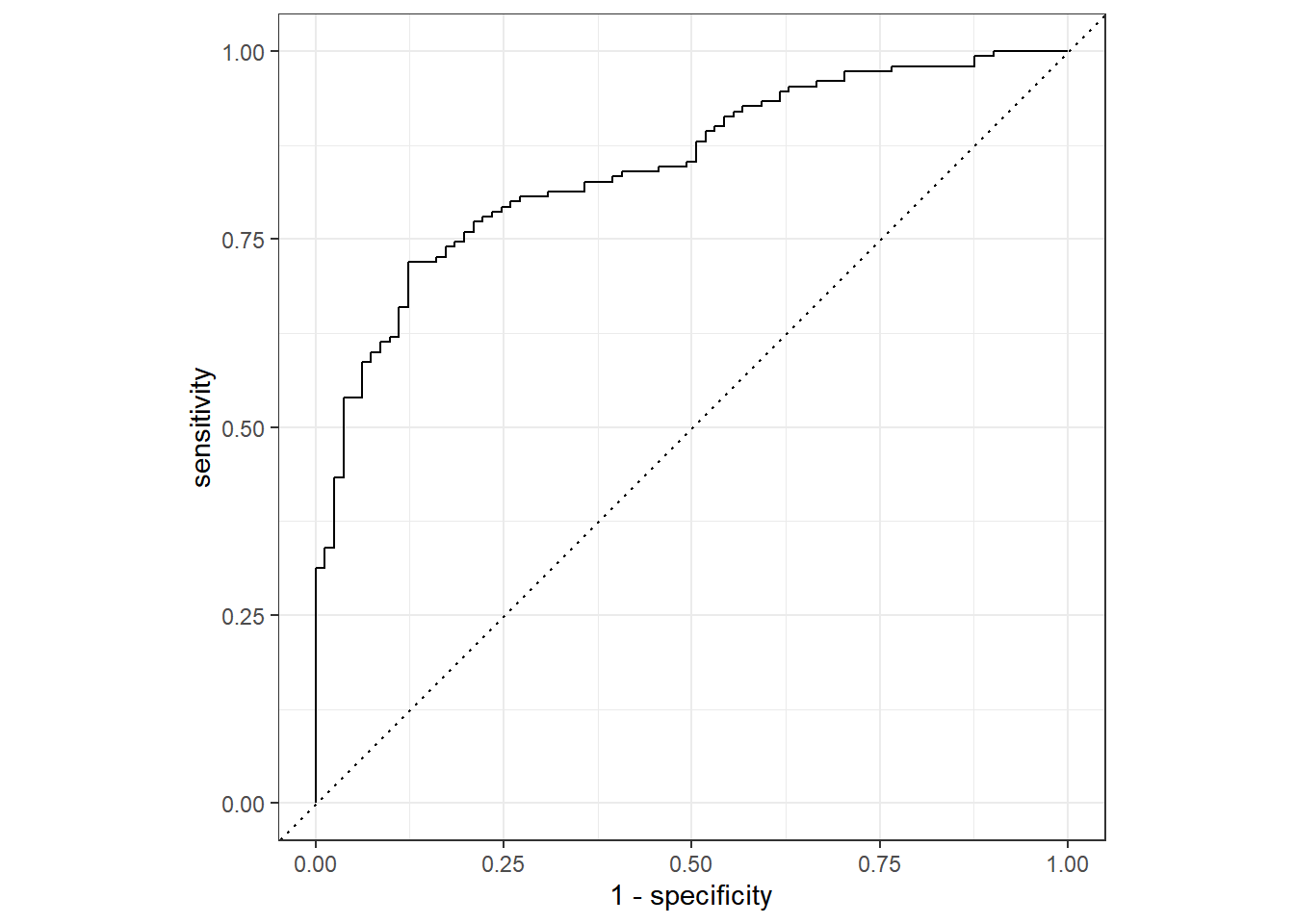
ranger_spec <- rand_forest(
mtry = tune(),
min_n = tune(),
trees = 1000
) %>%
set_mode("classification") %>%
set_engine("ranger", importance = "permutation")
ranger_recipe <-
recipe(formula = diabetes ~ ., data = pima_train)
ranger_wflow <- workflow() %>%
add_model(ranger_spec) %>%
add_recipe(ranger_recipe)
# 初始网格评估
set.seed(1234)
ranger_folds <- vfold_cv(pima_train)
ranger_params <- ranger_wflow %>%
extract_parameter_set_dials() %>%
finalize(pima_train)
ranger_params
## 设置初始网格
start_grid <-
ranger_params %>%
grid_regular(levels = 5)
## 初始网格搜索
ranger_initial <- ranger_wflow %>%
tune_grid(
resamples = ranger_folds,
grid = start_grid,
metrics = metric_set(roc_auc),
control = control_grid(save_pred = TRUE)
)
collect_metrics(ranger_initial)# A tibble: 25 × 8
mtry min_n .metric .estimator mean n std_err .config
<int> <int> <chr> <chr> <dbl> <int> <dbl> <chr>
1 1 2 roc_auc binary 0.839 10 0.0188 pre0_mod01_post0
2 1 11 roc_auc binary 0.845 10 0.0197 pre0_mod02_post0
3 1 21 roc_auc binary 0.847 10 0.0201 pre0_mod03_post0
4 1 30 roc_auc binary 0.845 10 0.0194 pre0_mod04_post0
5 1 40 roc_auc binary 0.847 10 0.0206 pre0_mod05_post0
6 3 2 roc_auc binary 0.839 10 0.0193 pre0_mod06_post0
7 3 11 roc_auc binary 0.842 10 0.0184 pre0_mod07_post0
8 3 21 roc_auc binary 0.844 10 0.0189 pre0_mod08_post0
9 3 30 roc_auc binary 0.846 10 0.0193 pre0_mod09_post0
10 3 40 roc_auc binary 0.847 10 0.0189 pre0_mod10_post0
# ℹ 15 more rows## 贝叶斯优化
ranger_tune_bayes <- ranger_wflow %>%
tune_bayes(
resamples = ranger_folds,
initial = ranger_initial, # 初始网格搜索结果
param_info = ranger_params,
iter = 25,
metrics = metric_set(roc_auc),
control = control_bayes(verbose = TRUE)
)
show_best(ranger_tune_bayes)# A tibble: 5 × 9
mtry min_n .metric .estimator mean n std_err .config .iter
<int> <int> <chr> <chr> <dbl> <int> <dbl> <chr> <int>
1 2 40 roc_auc binary 0.848 10 0.0204 iter01 1
2 2 35 roc_auc binary 0.848 10 0.0201 iter11 11
3 1 21 roc_auc binary 0.847 10 0.0201 pre0_mod03_post0 0
4 3 40 roc_auc binary 0.847 10 0.0189 pre0_mod10_post0 0
5 1 40 roc_auc binary 0.847 10 0.0206 pre0_mod05_post0 0# 拟合最终模型
## 选择最优超参数
ranger_best_bayes <- select_best(ranger_tune_bayes, metric = "roc_auc")
## 将最优参数应用至workflow
ranger_final_wflow <- ranger_wflow %>%
finalize_workflow(ranger_best_bayes)
ranger_final_wflow══ Workflow ════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: rand_forest()
── Preprocessor ────────────────────────────────────────────────────────────────
0 Recipe Steps
── Model ───────────────────────────────────────────────────────────────────────
Random Forest Model Specification (classification)
Main Arguments:
mtry = 2
trees = 1000
min_n = 40
Engine-Specific Arguments:
importance = permutation
Computational engine: ranger ## 拟合最终模型
ranger_final_fit <-
ranger_final_wflow %>%
last_fit(pima_split)
ranger_final_fit# Resampling results
# Manual resampling
# A tibble: 1 × 6
splits id .metrics .notes .predictions .workflow
<list> <chr> <list> <list> <list> <list>
1 <split [537/231]> train/test split <tibble> <tibble> <tibble> <workflow># 模型结果
ranger_final_fit %>%
collect_metrics()# A tibble: 3 × 4
.metric .estimator .estimate .config
<chr> <chr> <dbl> <chr>
1 accuracy binary 0.740 pre0_mod0_post0
2 roc_auc binary 0.843 pre0_mod0_post0
3 brier_class binary 0.153 pre0_mod0_post0# A tibble: 231 × 4
diabetes .pred_class .pred_pos .pred_neg
<fct> <fct> <dbl> <dbl>
1 neg neg 0.238 0.762
2 neg pos 0.529 0.471
3 pos pos 0.989 0.0106
4 neg neg 0.295 0.705
5 neg pos 0.575 0.425
6 neg neg 0.406 0.594
7 neg pos 0.628 0.372
8 pos pos 0.708 0.292
9 neg neg 0.257 0.743
10 pos pos 0.945 0.0553
# ℹ 221 more rows## 混淆矩阵
conf_mat(final_augment, truth = diabetes, estimate = .pred_class) Truth
Prediction pos neg
pos 125 35
neg 25 46## roc_auc
roc_auc(final_augment, truth = diabetes, .pred_pos)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 roc_auc binary 0.843## 多个指标一同输出
ranger_metrics <- metric_set(accuracy, mcc, f_meas)
ranger_metrics(final_augment, truth = diabetes, estimate = .pred_class)# A tibble: 3 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 accuracy binary 0.740
2 mcc binary 0.415
3 f_meas binary 0.806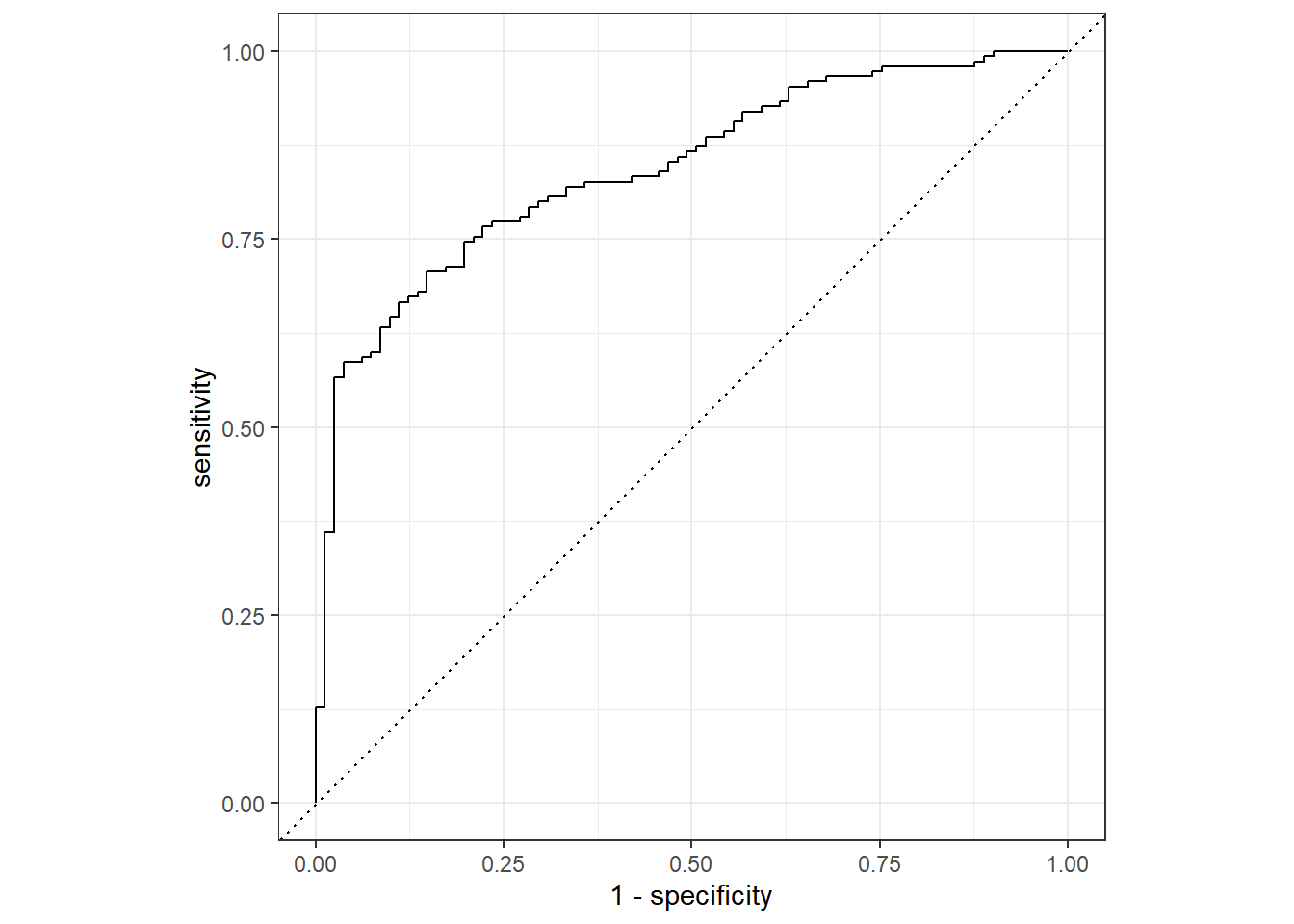
用网格搜索或迭代搜索对超参数进行调优后,模型结果都没有明显的提升。